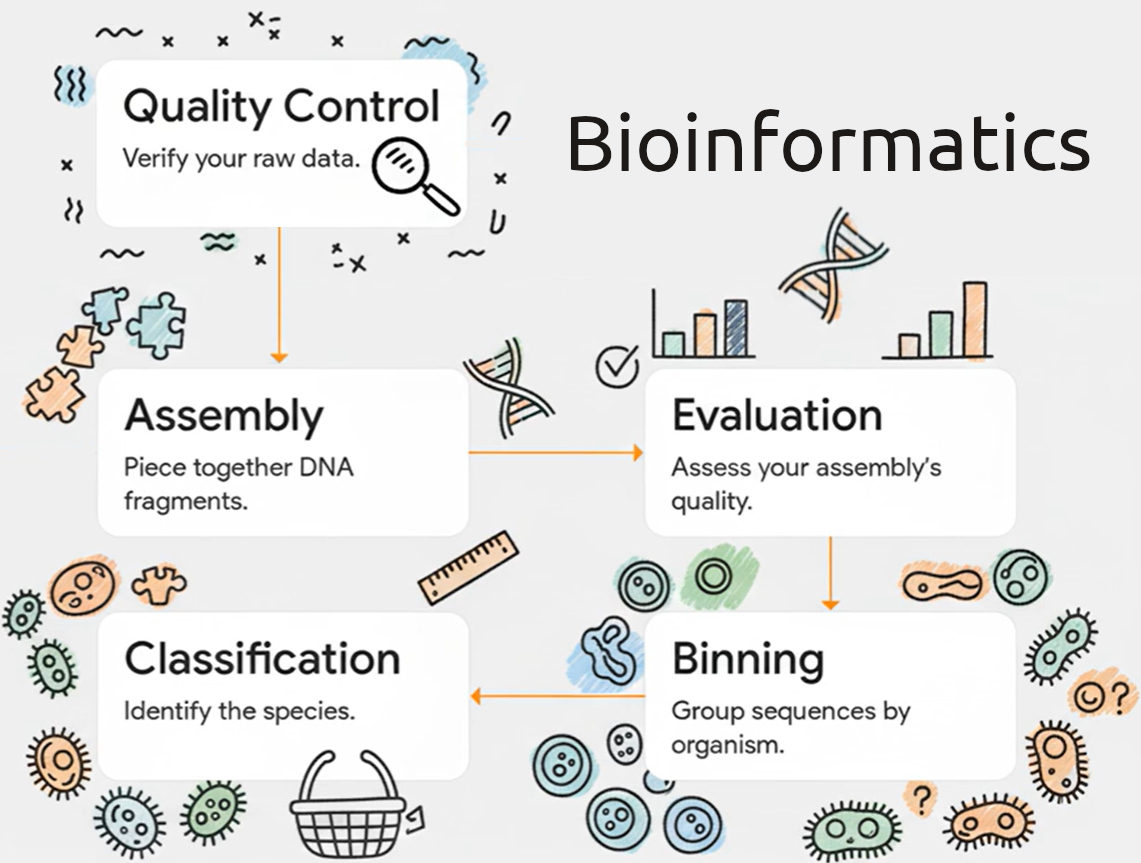
This self-paced course will explore a typical metagenomic pipeline to introduce some of the common tools used in bioinformatic analysis. The main objective of the course is to give you a head start in setting up a typical bioinformatic pipeline, like metagenomics, in an HPC environment rather than to be a complete course on the subject. As such, there is no mention of the identification of genes encoding for ribosomal RNAs (rRNAs), which are widely used for phylogenetic analysis and quantification of microbial diversity in a real-world metagenomic project.
The hands-on tutorial included in the course is a modified version of Alex Sczyrba’s Metagenomics Workshop to work with facilities provided by SHARCNET / the Alliance (formerly Compute Canada) and will guide you through the typical steps of metagenome assembly and binning. We will start with two FastQ files containing short paired-end reads of a simulated metagenome. Using a step-by-step metagenomics pipeline, we’re going to first assemble the whole metagenome using five different assemblers. Then, we will determine all the species in it (i.e. binning and classification).
The Certificate of Course Completion will be issued upon completion of the course and successful submission of the assignment at the end of it.
Prerequisites: none
Estimated time: 2 hours
- Teacher: Armin Sobhani
